Introduction to Geopandas¶
Downloading data¶
For this lesson we are using data in Shapefile format representing distributions of specific beautifully colored fish species called Damselfish and the country borders of Europe. From now on, we are going to download the datafiles at the start of each lesson because of the large size of the data. It is also a good practice to know how to download files from terminal.
On Binder and CSC Notebook environment, you can use wget programn to
download the data. Let’s download the
data into
folder /home/jovyan/notebooks/L2 by running following commands in
the Terminal (see
here
if you don’t know how to launch a terminal):
# Change directory to directory with Lesson 2 materials
$ cd /home/jovyan/notebooks/L2
$ wget https://github.com/AutoGIS/data/raw/master/L2_data.zip
Hint: you can copy/paste things to JupyterLab Terminal by pressing SHIFT + RIGHT-CLICK on your mouse and choosing ‘Paste’.
Once you have downloaded the L2_data.zip file into your home
directory, you can unzip the file using unzip command from Terminal
(or e.g. 7zip on Windows if working with own computer). Following
assumes that the file was downloaded to /home/jovyan/notebooks/L2
-directory:
$ cd /home/jovyan/notebooks/L2
$ unzip L2_data.zip
$ ls L2_data
DAMSELFISH_distributions.cpg DAMSELFISH_distributions.shp Europe_borders.dbf Europe_borders.sbx
DAMSELFISH_distributions.dbf DAMSELFISH_distributions.shx Europe_borders.prj Europe_borders.shp
DAMSELFISH_distributions.prj Europe_borders.cpg Europe_borders.sbn Europe_borders.shx
As we can see, the L2_data folder includes Shapefiles called
DAMSELFISH_distribution.shp and Europe_borders.shp. Notice that
Shapefile -fileformat is constituted of many separate files such as
.dbf that contains the attribute information, and .prj -file
that contains information about coordinate reference system.
Reading a Shapefile¶
Typically reading the data into Python is the first step of the analysis
pipeline. In GIS, there exists various dataformats such as
Shapefile,
GeoJSON,
KML, and
GPKG that are probably
the most common vector data formats.
Geopandas is capable of reading data
from all of these formats (plus many more). Reading spatial data can be
done easily with geopandas using gpd.from_file() -function:
In [2]:
# Import necessary modules
import geopandas as gpd
# Set filepath
fp = "L2_data/DAMSELFISH_distributions.shp"
# Read file using gpd.read_file()
data = gpd.read_file(fp)
Now we read the data from a Shapefile into variable data.
- Let’s see check the data type of it
In [3]:
type(data)
Out[3]:
geopandas.geodataframe.GeoDataFrame
Okey so from the above we can see that our data -variable is a
GeoDataFrame. GeoDataFrame extends the functionalities of
pandas.DataFrame in a way that it is possible to use and handle
spatial data using similar approaches and datastructures as in Pandas
(hence the name geopandas). GeoDataFrame have some special features and
functions that are useful in GIS.
- Let’s take a look at our data and print the first 2 rows using the
head()-function:
In [4]:
print(data.head(2))
ID_NO BINOMIAL ORIGIN COMPILER YEAR \
0 183963.0 Stegastes leucorus 1 IUCN 2010
1 183963.0 Stegastes leucorus 1 IUCN 2010
CITATION SOURCE DIST_COMM ISLAND \
0 International Union for Conservation of Nature... None None None
1 International Union for Conservation of Nature... None None None
SUBSPECIES ... RL_UPDATE \
0 None ... 2012.1
1 None ... 2012.1
KINGDOM_NA PHYLUM_NAM CLASS_NAME ORDER_NAME FAMILY_NAM \
0 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
1 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
GENUS_NAME SPECIES_NA CATEGORY \
0 Stegastes leucorus VU
1 Stegastes leucorus VU
geometry
0 POLYGON ((-115.6437454219999 29.71392059300007...
1 POLYGON ((-105.589950704 21.89339825500002, -1...
[2 rows x 24 columns]
As we can see, there exists multiple columns in our data related to our Damselfish -fish.
When having spatial data, it is always a good idea to explore your data
on a map. Creating a simple map from a GeoDataFrame is really easy:
you can use .plot() -function from geopandas that creates a map
based on the geometries of the data. Geopandas actually uses Matplotlib
for creating the map that was introduced in Lesson 7 of Geo-Python
course.
- Let’s try it out, and take a look how our data looks like on a map:
In [7]:
%matplotlib inline
data.plot()
Out[7]:
<matplotlib.axes._subplots.AxesSubplot at 0x18feb9164a8>
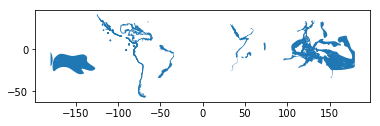
Voilá! As we can see, it is really easy to produce a map out of your Shapefile with geopandas. Geopandas automatically positions your map in a way that it covers the whole extent of your data.
Writing a Shapefile¶
Writing the spatial data into disk for example as a new Shapefile is also something that is needed frequently.
- Let’s select 50 first rows of the input data and write those into a
new Shapefile by first selecting the data using index slicing and
then write the selection into a Shapefile with
gpd.to_file()-function:
In [8]:
# Create a output path for the data
outfp = "L2_data/DAMSELFISH_distributions_SELECTION.shp"
# Select first 50 rows
selection = data[0:50]
# Write those rows into a new Shapefile (the default output file format is Shapefile)
selection.to_file(outfp)
TASK: Read the newly created Shapefile with geopandas, and see how the data looks like.
Geometries in Geopandas¶
Geopandas takes advantage of Shapely’s geometric objects. Geometries are stored in a column called geometry that is a default column name for storing geometric information in geopandas.
- Let’s print the first 5 rows of the column ‘geometry’:
In [10]:
# It is possible to get a specific column by specifying the column name within square brackets []
print(data['geometry'].head())
0 POLYGON ((-115.6437454219999 29.71392059300007...
1 POLYGON ((-105.589950704 21.89339825500002, -1...
2 POLYGON ((-111.159618439 19.01535626700007, -1...
3 POLYGON ((-80.86500229899997 -0.77894492099994...
4 POLYGON ((-67.33922225599997 -55.6761029239999...
Name: geometry, dtype: object
As we can see the geometry column contains familiar looking values,
namely Shapely Polygon -objects that we learned to use last
week.
Since the spatial data is stored as Shapely objects, it is possible to
use all of the functionalities of Shapely module.
- Let’s prove that this really is the case by iterating over a sample
of the data, and printing the
areaof first five polygons.- We can iterate over the rows by using the
iterrows()-function that we learned during the Lesson 6 of the Geo-Python course.
- We can iterate over the rows by using the
In [11]:
# Make a selection that contains only the first five rows
selection = data[0:5]
# Iterate over rows and print the area of a Polygon
for index, row in selection.iterrows():
# Get the area of the polygon
poly_area = row['geometry'].area
# Print information for the user
print("Polygon area at index {index} is: {area:.3f}".format(index=index, area=poly_area))
Polygon area at index 0 is: 19.396
Polygon area at index 1 is: 6.146
Polygon area at index 2 is: 2.697
Polygon area at index 3 is: 87.461
Polygon area at index 4 is: 0.001
As you might guess from here, all the functionalities of Pandas,
such as the iterrows() function, are directly available in Geopandas
without the need to call pandas separately because Geopandas is an
extension for Pandas.
- Let’s next create a new column into our GeoDataFrame where we
calculate and store the areas of individual polygons into that
column. Calculating the areas of polygons is really easy in geopandas
by using
GeoDataFrame.areaattribute. Hence, it is not needed to actually iterate over the rows line by line as we did previously:
In [14]:
# Create a new column called 'area' and assign the area of the Polygons into it
data['area'] = data.area
# Print first 2 rows of the area column
print(data['area'].head(2))
0 19.396254
1 6.145902
Name: area, dtype: float64
As we can see, the area of our first polygon seems to be approximately
19.396 and 6.146 for the second polygon. They correspond to the
ones we saw in previous step when iterating rows, hence, everything
seems to work as should.
- Let’s check what is the
min,maxandmeanof those areas using familiar functions from our previous Pandas lessions.
In [16]:
# Maximum area
max_area = data['area'].max()
# Minimum area
min_area = data['area'].min()
# Mean area
mean_area = data['area'].mean()
print("Max area: {max}\nMin area: {min}\nMean area: {mean}".format(max=round(max_area, 2), min=round(min_area, 2), mean=round(mean_area, 2)))
Max area: 1493.2
Min area: 0.0
Mean area: 19.96
The largest Polygon in our dataset seems to be around 1494 square
decimal degrees (~ 165 000 km2) and the average size is ~20 square
decimal degrees (~2200 km2). The minimum polygon size seems to be
0.0, hence it seems that there exists really small polygons as well
in the data as well (rounds to 0 with 2 decimals).
Creating geometries into a GeoDataFrame¶
Since geopandas takes advantage of Shapely geometric objects, it is possible to create a Shapefile from a scratch by passing Shapely’s geometric objects into the GeoDataFrame. This is useful as it makes it easy to convert e.g. a text file that contains coordinates into a Shapefile. Next we will see how to create a Shapefile from scratch.
- Let’s create an empty
GeoDataFrame.
In [28]:
# Import necessary modules first
import geopandas as gpd
from shapely.geometry import Point, Polygon
# Create an empty geopandas GeoDataFrame
newdata = gpd.GeoDataFrame()
# Let's see what we have at the moment
print(newdata)
Empty GeoDataFrame
Columns: []
Index: []
As we can see, the GeoDataFrame is empty since we haven’t yet stored any data into it.
- Let’s create a new column called
geometrythat will contain our Shapely objects:
In [29]:
# Create a new column called 'geometry' to the GeoDataFrame
newdata['geometry'] = None
# Let's again see what's inside
print(newdata)
Empty GeoDataFrame
Columns: [geometry]
Index: []
Now we have a geometry column in our GeoDataFrame but we don’t have
any data stored yet.
- Let’s create a Shapely
Polygonrepsenting the Helsinki Senate square that we can later insert to our GeoDataFrame:
In [30]:
# Coordinates of the Helsinki Senate square in Decimal Degrees
coordinates = [(24.950899, 60.169158), (24.953492, 60.169158), (24.953510, 60.170104), (24.950958, 60.169990)]
# Create a Shapely polygon from the coordinate-tuple list
poly = Polygon(coordinates)
# Let's see what we have
print(poly)
POLYGON ((24.950899 60.169158, 24.953492 60.169158, 24.95351 60.170104, 24.950958 60.16999, 24.950899 60.169158))
Okay, now we have an appropriate Polygon -object.
- Let’s insert the polygon into our ‘geometry’ column of our GeoDataFrame at position 0:
In [31]:
# Insert the polygon into 'geometry' -column at index 0
newdata.loc[0, 'geometry'] = poly
# Let's see what we have now
print(newdata)
geometry
0 POLYGON ((24.950899 60.169158, 24.953492 60.16...
Great, now we have a GeoDataFrame with a Polygon that we could already now export to a Shapefile. However, typically you might want to include some useful information with your geometry.
- Hence, let’s add another column to our GeoDataFrame called
locationwith textSenaatintorithat describes the location of the feature.
In [32]:
# Add a new column and insert data
newdata.loc[0, 'location'] = 'Senaatintori'
# Let's check the data
print(newdata)
geometry location
0 POLYGON ((24.950899 60.169158, 24.953492 60.16... Senaatintori
Okay, now we have additional information that is useful for recognicing what the feature represents.
Before exporting the data it is always good (basically necessary) to
determine the coordinate reference system (projection) for the
GeoDataFrame. GeoDataFrame has an attribute called .crs that shows
the coordinate system of the data which is empty (None) in our case
since we are creating the data from the scratch (more about projection
on next tutorial):
In [33]:
print(newdata.crs)
None
- Let’s add a crs for our GeoDataFrame. A Python module called
fiona has a nice function called
from_epsg()for passing the coordinate reference system information for the GeoDataFrame. Next we will use that and determine the projection to WGS84 (epsg code: 4326):
In [34]:
# Import specific function 'from_epsg' from fiona module
from fiona.crs import from_epsg
# Set the GeoDataFrame's coordinate system to WGS84 (i.e. epsg code 4326)
newdata.crs = from_epsg(4326)
# Let's see how the crs definition looks like
print(newdata.crs)
{'init': 'epsg:4326', 'no_defs': True}
As we can see, now we have associated coordinate reference system
information (i.e. CRS) into our GeoDataFrame. The CRS
information here, is a Python dictionary containing necessary values
for geopandas to create a .prj file for our Shapefile that contains
the CRS info.
- Finally, we can export the GeoDataFrame using
.to_file()-function. The function works quite similarly as the export functions in numpy or pandas, but here we only need to provide the output path for the Shapefile. Easy isn’t it!:
In [35]:
# Determine the output path for the Shapefile
outfp = "L2_data/Senaatintori.shp"
# Write the data into that Shapefile
newdata.to_file(outfp)
Now we have successfully created a Shapefile from the scratch using only Python programming. Similar approach can be used to for example to read coordinates from a text file (e.g. points) and create Shapefiles from those automatically.
TASK: Check the output Shapefile by reading it with geopandas and make sure that the attribute table and geometry seems correct.
Practical example: Saving multiple Shapefiles¶
One really useful function that can be used in Pandas/Geopandas is .groupby(). We saw and used this function already in Lesson 6 of the Geo-Python course. Group by function is useful to group data based on values on selected column(s).
Next we will take a practical example by automating the file export
task. We will group individual fish subspecies in our
DAMSELFISH_distribution.shp and export those into separate
Shapefiles.
- Let’s start from scratch and read the Shapefile into GeoDataFrame
In [37]:
# Read Damselfish data
fp = "L2_data/DAMSELFISH_distributions.shp"
data = gpd.read_file(fp)
# Print columns
print(data.columns)
Index(['ID_NO', 'BINOMIAL', 'ORIGIN', 'COMPILER', 'YEAR', 'CITATION', 'SOURCE',
'DIST_COMM', 'ISLAND', 'SUBSPECIES', 'SUBPOP', 'LEGEND', 'SEASONAL',
'TAX_COMM', 'RL_UPDATE', 'KINGDOM_NA', 'PHYLUM_NAM', 'CLASS_NAME',
'ORDER_NAME', 'FAMILY_NAM', 'GENUS_NAME', 'SPECIES_NA', 'CATEGORY',
'geometry'],
dtype='object')
The BINOMIAL column in the data contains information about different
fish subspecies (their latin name). With .unique() -function we can
quickly see all different names in that column:
In [38]:
# Print all unique fish subspecies in 'BINOMIAL' column
print(data['BINOMIAL'].unique())
['Stegastes leucorus' 'Chromis intercrusma' 'Stegastes beebei'
'Stegastes rectifraenum' 'Chromis punctipinnis' 'Chromis crusma'
'Chromis pembae' 'Stegastes redemptus' 'Teixeirichthys jordani'
'Chromis limbaughi' 'Microspathodon dorsalis' 'Chromis cyanea'
'Amphiprion sandaracinos' 'Nexilosus latifrons' 'Stegastes baldwini'
'Microspathodon bairdii' 'Azurina eupalama' 'Chromis flavicauda'
'Stegastes arcifrons' 'Chromis alta' 'Abudefduf declivifrons'
'Chromis alpha' 'Stegastes flavilatus' 'Abudefduf concolor'
'Abudefduf troschelii' 'Chrysiptera flavipinnis' 'Chromis atrilobata'
'Stegastes acapulcoensis' 'Hypsypops rubicundus' 'Azurina hirundo']
- Now we can use that information to group our data and save all individual fish subspecies as separate Shapefiles:
In [39]:
# Group the data by column 'BINOMIAL'
grouped = data.groupby('BINOMIAL')
# Let's see what we have
grouped
Out[39]:
<pandas.core.groupby.groupby.DataFrameGroupBy object at 0x0000018FF1F87208>
As we can see, groupby -function gives us an object called
DataFrameGroupBy which is similar to list of keys and values (in a
dictionary) that we can iterate over. This is again exactly similar
thing that we already practiced during Lesson 6 of the Geo-Python
course.
- Let’s iterate over the groups and see what our variables
keyandvaluescontain
In [41]:
# Iterate over the group object
for key, values in grouped:
individual_fish = values
# Let's see what is the LAST item and key that we iterated
print('Key:', key)
print(individual_fish)
Key: Teixeirichthys jordani
ID_NO BINOMIAL ORIGIN COMPILER YEAR \
27 154915.0 Teixeirichthys jordani 1 None 2012
28 154915.0 Teixeirichthys jordani 1 None 2012
29 154915.0 Teixeirichthys jordani 1 None 2012
30 154915.0 Teixeirichthys jordani 1 None 2012
31 154915.0 Teixeirichthys jordani 1 None 2012
32 154915.0 Teixeirichthys jordani 1 None 2012
33 154915.0 Teixeirichthys jordani 1 None 2012
CITATION SOURCE DIST_COMM ISLAND \
27 Red List Index (Sampled Approach), Zoological ... None None None
28 Red List Index (Sampled Approach), Zoological ... None None None
29 Red List Index (Sampled Approach), Zoological ... None None None
30 Red List Index (Sampled Approach), Zoological ... None None None
31 Red List Index (Sampled Approach), Zoological ... None None None
32 Red List Index (Sampled Approach), Zoological ... None None None
33 Red List Index (Sampled Approach), Zoological ... None None None
SUBSPECIES ... RL_UPDATE \
27 None ... 2012.2
28 None ... 2012.2
29 None ... 2012.2
30 None ... 2012.2
31 None ... 2012.2
32 None ... 2012.2
33 None ... 2012.2
KINGDOM_NA PHYLUM_NAM CLASS_NAME ORDER_NAME FAMILY_NAM \
27 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
28 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
29 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
30 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
31 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
32 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
33 ANIMALIA CHORDATA ACTINOPTERYGII PERCIFORMES POMACENTRIDAE
GENUS_NAME SPECIES_NA CATEGORY \
27 Teixeirichthys jordani LC
28 Teixeirichthys jordani LC
29 Teixeirichthys jordani LC
30 Teixeirichthys jordani LC
31 Teixeirichthys jordani LC
32 Teixeirichthys jordani LC
33 Teixeirichthys jordani LC
geometry
27 POLYGON ((121.6300326400001 33.04248618400004,...
28 POLYGON ((32.56219482400007 29.97488975500005,...
29 POLYGON ((130.9052090560001 34.02498196400006,...
30 POLYGON ((56.32233070000007 -3.707270205999976...
31 POLYGON ((40.64476131800006 -10.85502363999996...
32 POLYGON ((48.11258402900006 -9.335103113999935...
33 POLYGON ((51.75403543100003 -9.21679305899994,...
[7 rows x 24 columns]
From here we can see that the individual_fish -variable contains all
the rows that belongs to a fish called Teixeirichthys jordani that
is the key for conducting the grouping. Notice that the index
numbers refer to the row numbers in the original data -GeoDataFrame.
- Let’s check the datatype of the grouped object:
In [42]:
type(individual_fish)
Out[42]:
geopandas.geodataframe.GeoDataFrame
As we can see, each set of data are now grouped into separate
GeoDataFrames that we can export into Shapefiles using the variable
key for creating the output filename. Next, we use a specific string
formatting method to produce the output filename using % operator
(read more here).
- Let’s now export all individual subspecies into separate Shapefiles:
In [45]:
# Import os -module that is useful for parsing filepaths
import os
# Determine output directory
out_directory = "L2_data"
# Create a new folder called 'Results'
result_folder = os.path.join(out_directory, 'Results')
# Check if the folder exists already
if not os.path.exists(result_folder):
# If it does not exist, create one
os.makedirs(result_folder)
# Iterate over the groups
for key, values in grouped:
# Format the filename (replace spaces with underscores using 'replace()' -function)
output_name = "%s.shp" % key.replace(" ", "_")
# Print some information for the user
print("Processing: %s" % key)
# Create an output path
outpath = os.path.join(result_folder, output_name)
# Export the data
values.to_file(outpath)
Processing: Abudefduf concolor
Processing: Abudefduf declivifrons
Processing: Abudefduf troschelii
Processing: Amphiprion sandaracinos
Processing: Azurina eupalama
Processing: Azurina hirundo
Processing: Chromis alpha
Processing: Chromis alta
Processing: Chromis atrilobata
Processing: Chromis crusma
Processing: Chromis cyanea
Processing: Chromis flavicauda
Processing: Chromis intercrusma
Processing: Chromis limbaughi
Processing: Chromis pembae
Processing: Chromis punctipinnis
Processing: Chrysiptera flavipinnis
Processing: Hypsypops rubicundus
Processing: Microspathodon bairdii
Processing: Microspathodon dorsalis
Processing: Nexilosus latifrons
Processing: Stegastes acapulcoensis
Processing: Stegastes arcifrons
Processing: Stegastes baldwini
Processing: Stegastes beebei
Processing: Stegastes flavilatus
Processing: Stegastes leucorus
Processing: Stegastes rectifraenum
Processing: Stegastes redemptus
Processing: Teixeirichthys jordani
Excellent! Now we have saved those individual fishes into separate Shapefiles and named the file according to the species name. These kind of grouping operations can be really handy when dealing with Shapefiles. Doing similar process manually would be really laborious and error-prone.
Summary¶
In this tutorial we introduced the first steps of using geopandas. More specifically you should know how to:
1) Read data from Shapefile using geopandas,
2) Write GeoDataFrame data from Shapefile using geopandas,
3) Create a GeoDataFrame from scratch, and
4) automate a task to save specific rows from data into Shapefile
based on specific key using groupby() -function.

